
Building Your First R Package: Day 3
In our final session, we will wrap up our package development and learn how to share our tools with the world by publishing to GitHub and to CRAN.
0.1 Introduction: From Local to Global
Welcome to Day 3! Over the last two days, we transitioned from script-writers to package-developers. We built the vizwizard package, packed it with custom ggplot2 and gt wrappers, and learned how to bundle our own custom datasets.
To reiterate, effective package design can be summarized in the below diagram/flowchart. Whether you’re leading a classroom instruction and want to create a package for students to share datasets, functions, etc. or spreading your love for medical analysis and alleviating headaches for researchers/physicians, the below chart is helpful.
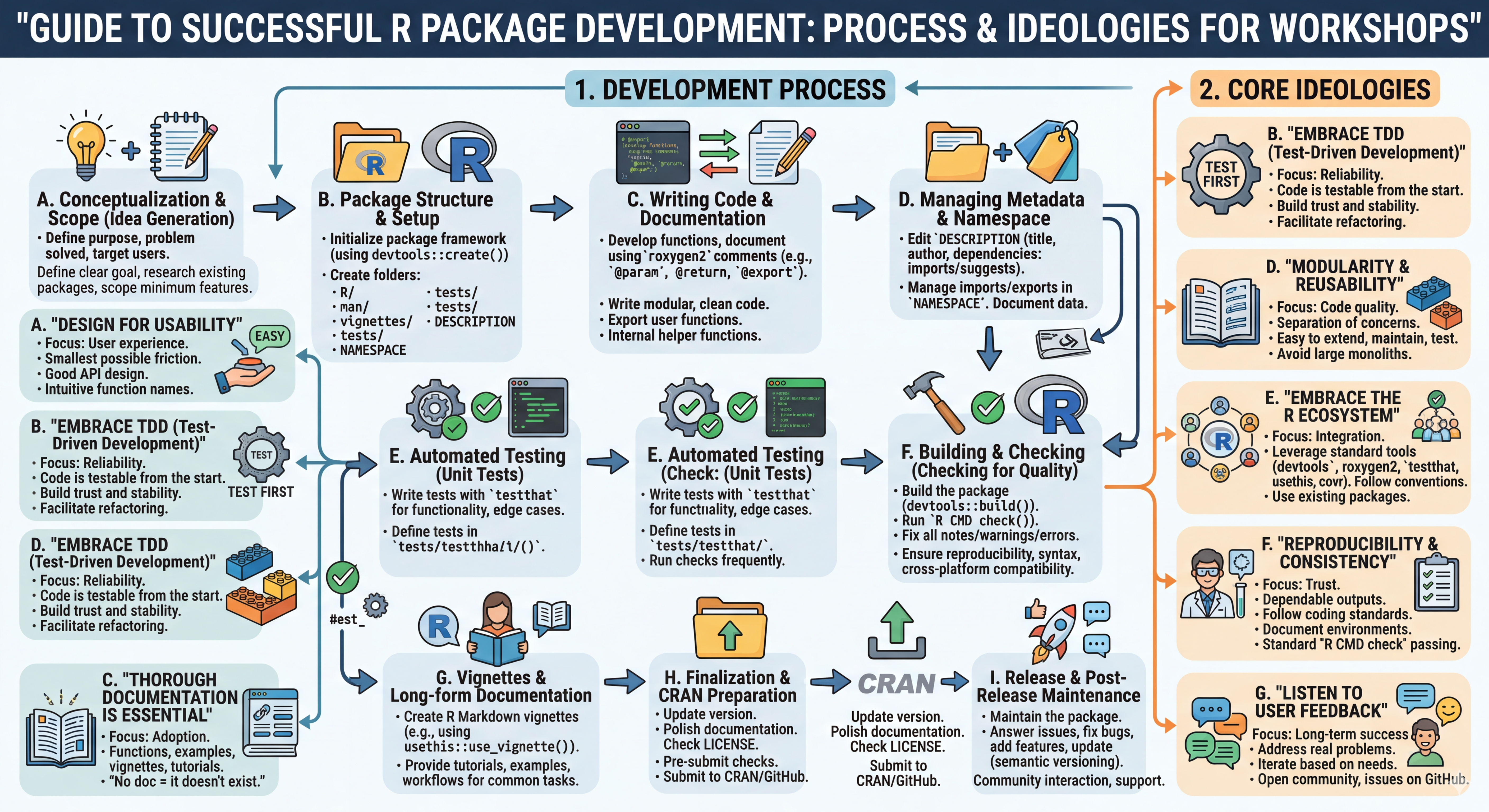
Today is all about deployment. An R package is incredibly useful for your own reproducibility, but its true power is unlocked when you share it.
Today we will cover:
- Tying Up Loose Ends: Making sure our package is completely finished and error-free.
- Publishing to GitHub (From Scratch): The modern, fast standard for sharing packages with colleagues, including how to authenticate your account.
- Publishing to CRAN: Understanding the strict rules and rigorous checks required for the official R repository.
- Troubleshooting: Solving common deployment problems.
Let’s load our tools one last time!
library(devtools)Loading required package: usethislibrary(usethis)1 Part 1: Tying Up Loose Ends
Before we can publish anything, we need to ensure our house is in order.
1.1 The Final Check
On Day 2, we learned about the ultimate package spellcheck: devtools::check(). This function is the gatekeeper for publishing.
Run it now in your console:
devtools::check()Look at the bottom of your console output. Your goal before proceeding is 0 Errors, 0 Warnings, and 0 Notes!
Errors: Your package is fundamentally broken. Must be fixed.
Warnings: Your package works, but you did something risky. Must be fixed if you want to go to CRAN.
Notes: Minor formatting suggestions. CRAN reviewers might let a Note slide, but it’s best practice to resolve them.
A common error from previous weeks surrounds dependency errors - when a package is needed but not called/referenced. If this occurs, ensure that there are no function calls in your R scripts and that every script declares 1 function! See the previous trouble shooting tip from other sessions:
Remember, while you can/should put the pubscatter() command in the same file as the function() for evaluation purposes (to test if your code works), remember to delete this line before you build/compile/send your package out for others to use.
2 Part 2: Publishing to GitHub (From Scratch)
CRAN is great, but it has extremely strict rules. If you want to share a package with your lab mates immediately, or if you are hosting internal clinical code that shouldn’t be public, GitHub is the way to go.
2.1 Step 1: Version Control with Git
Your project needs to be a Git repository before it can go to GitHub. The usethis package makes this a one-liner. Run this in your console:
usethis::use_git()RStudio will ask if it is okay to commit your files and restart. Select “Yes” to both.
2.2 Step 2: Authentication (The Tricky Part!)
To send code from your local computer to GitHub, GitHub needs to know it’s really you. We do this using a Personal Access Token.
- Create the Token: Run this command. It will open a GitHub webpage in your browser.
usethis::create_github_token()Scroll down on the webpage and click “Generate Token”.
Copy the token (it looks like a long string of random letters and numbers). Do not lose this!
Give the Token to R: Go back to RStudio and run:
Paste your token into the console when prompted and hit Enter.
2.3 Step 3: Push to GitHub
Now that RStudio and GitHub are linked, you can push your package to the cloud!
usethis::use_github()How users install your GitHub package Once your package is on GitHub, you can tell your colleagues to run this code to download and install your tools:
# They only need to run this once
install.packages("remotes")
remotes::install_github("your-username/vizwizard")3 Part 3: Publishing to CRAN
The Comprehensive R Archive Network (CRAN) is the official repository for R packages. Publishing here means your package can be installed with a simple install.packages("vizwizard").
Understanding CRAN Rules and Requirements
CRAN is notoriously strict. They are protecting the entire R ecosystem from malicious code, broken dependencies, and bloated file sizes. Here is what CRAN strictly requires:
Zero Errors or Warnings: Your R CMD check must pass cleanly on Windows, Mac, Linux, and Solaris.
File Size Limits: Your package must generally be under 5MB. If you include massive datasets, you will be rejected.
Impeccable Documentation: Every single exported function must have a complete help file with working examples. If an example throws an error, the package is rejected.
Standard Licenses: You must clearly declare an open-source license (e.g., MIT or GPL-3) in your DESCRIPTION file.
No “Hidden” Changes: Your package cannot modify the user’s global .Rprofile, secretly install other software, or change their working directory without permission.
3.1 Step 1: Cross-Platform Testing
Your computer is only one operating system. R has a built-in “inspector” that checks your package, but CRAN requires your package to work on all operating systems. devtools provides ways to send your package to external servers to be tested.
Run these and wait for the email results:
# Test on a CRAN-like Windows server
devtools::check_win_devel()
# Test on a CRAN-like Mac server
devtools::check_mac_release()3.2 Step 2: The Final Submission
If your local devtools::check() is totally clean, and your cross-platform checks came back green, you are ready to submit!
devtools::release()R will ask you a series of questions to double-check that you are absolutely ready. Once you say yes, it bundles your package and emails it to the CRAN team. You will usually hear back from a volunteer reviewer within a few days.
4 Part 4: Practice Problems & Troubleshooting
Package development rarely goes perfectly on the first try. Let’s walk through some common problems.
4.1 Problem 1: The DESCRIPTION File Dependency Error
Scenario: You ran devtools::check() and got the following Error:
Error: namespace ‘ggplot2’ is not available and has been NOTED.
Click for the solution
Why it happened: You wrote a function that relies on ggplot2, but you forgot to tell your package that it requires it! R doesn’t know to load it.
The Fix: As we learned on Day 1, never type package names into the DESCRIPTION file manually. Run this in your console to automatically add it:
usethis::use_package("ggplot2")
Bonus: What if the problem was missing dplyr or gt?
The fix is exactly the same! Just replace ggplot2 with the missing package name.
4.2 Problem 2: The “Undocumented Argument” Warning
Scenario: You ran devtools::check() and received a Warning:
Warning: Undocumented arguments in documentation object ‘pub_scatter’: ‘color_var’
Click for the solution
Why it happened: You added a new argument (color_var) to your custom scatterplot function, but you forgot to write a roxygen2 comment for it. CRAN will reject this immediately because the help file doesn’t match the function.
The Fix:
Open your R/plots.R script.
Find the pub_scatter function.
Add the missing tag above the function: #’ @param color_var The variable to color points by (unquoted)
Run devtools::document() to update the manual.
Re-run devtools::check().
4.3 Problem 3: The CRAN Rejection
Scenario: You submitted to CRAN, but two days later you get an email from a reviewer saying:
“Please do not use T and F in your code. Spell out TRUE and FALSE. Please fix and resubmit.”
Click for the solution
Why it happened: In R, T and F are technically variables that can be overwritten by the user (e.g., someone could run T <- FALSE in their console and break everything). CRAN prohibits the use of T and F abbreviations in package code.
The Fix:
Use Ctrl+F (or Cmd+F) to search your entire project for instances of T and F.
Change them to TRUE and FALSE.
Open your DESCRIPTION file and bump your version number up (e.g., change 0.1.0 to 0.1.1).
Re-run devtools::check() to ensure everything is perfect.
Submit again using devtools::release().
Conclusion: You Did It! Congratulations! Over the last three days, you have conquered one of the steepest learning curves in the R ecosystem. You’ve learned how to harness usethis to generate boilerplate code , use tidy evaluation ({{ }}) for custom functions , bundle data, and ultimately share your work with the world. +2
Whether you keep your packages local, put them on GitHub, or send them to CRAN, you are now an R Package Developer. Happy coding!