# 1 - Introduction ----------------------------------------------------
# 1.4 - Loading the tidyverse
# 2 - The gapminder dataset ------------------------------------------
# 3 - arrange() -------------------------------------------------------
# Challenge 1
# 4 - filter() --------------------------------------------------------
# 4.1 - Filter based on one condition
# 4.2 - Filter based on multiple conditions
# 5 - The pipe --------------------------------------------------------
# Challenge 2
# Challenge 3
# 6 - mutate() --------------------------------------------------------
# Challenge 4
# 7 - summarize() -----------------------------------------------------
# Challenge 5Data wrangling with dplyr
1 Introduction
1.1 What we’ll cover
Many of the most common data analysis tasks involve working with tabular data, i.e. data with rows and columns. In this session, you’ll learn to work with tabular data in R using the dplyr package.
For example, you’ll see how to subset, manipulate and summarize a dataset, the kinds of tasks commonly referred to as “data wrangling”. Such data processing is often essential before you can move on to say, data visualization (next session of this workshop) or statistical analyses.
1.2 Setting up
Use a new script for this session — much like in the previous session:
Open a new R script (Click the
+symbol in toolbar at the top, then clickR Script)1.Save the script straight away as
data-wrangling.R– you can save it anywhere you like, though it is probably best to save it in a folder specifically for this workshop.If you want the section headers as comments in your script, like in the script I am showing you in the live session, then copy-and-paste the following into your script:
Section headers for your script (Click to expand)
1.3 The dplyr package and the tidyverse
One of R’s most powerful features is its ability to deal with tabular data: data with rows and columns like you are familiar with from Excel spreadsheets and so on – for example:
country continent year lifeExp pop gdpPercap
1 Afghanistan Asia 1952 28.801 8425333 779.4453
2 Afghanistan Asia 1957 30.332 9240934 820.8530
3 Afghanistan Asia 1962 31.997 10267083 853.1007
4 Afghanistan Asia 1967 34.020 11537966 836.1971
5 Afghanistan Asia 1972 36.088 13079460 739.9811
6 Afghanistan Asia 1977 38.438 14880372 786.1134In R, tabular data is stored in a data structure called “data frame”. The dplyr package provides very useful functions for manipulating data in data frames. In this session, we’ll cover these commonly used dplyr functions:
arrange()to change the order of rows (i.e., to sort a data frame)filter()to keep only a subset of rowsmutate()to manipulate columns and create new columnssummarize()to compute data summaries across rows
dplyr has many more functions. Some of these are more advanced than those we’ll cover today, while others are conceptually simpler (e.g. rename() to change column names and select() to select columns) and we encourage you to explore these on your own.
dplyr belongs to a family of R packages for data science called the &&“tidyverse”.** Another tidyverse package we’ll cover in today’s workshop is ggplot2 for making plots.
1.4 Loading the tidyverse
All core tidyverse packages can be installed and loaded with a single command. Since you should already have installed the tidyverse2, you only need to load it, which you do as follows:
library(tidyverse)── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.4 ✔ readr 2.1.5
✔ forcats 1.0.1 ✔ stringr 1.5.2
✔ ggplot2 4.0.0 ✔ tibble 3.3.0
✔ lubridate 1.9.4 ✔ tidyr 1.3.1
✔ purrr 1.1.0
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorsThe output tells you which packages have been loaded as part of the tidyverse. (For now, you don’t have to worry about the “Conflicts” section.)
2 The gapminder dataset
In this session and the next one on data visualization, we will work with the gapminder dataset, which contains statistics such as population size for different countries across five-year intervals.
This dataset is available in a package of the same name, which you already should have installed as well3. To load the package into your current R session:
# (Unlike with the tidyverse, no output is expected when you load gapminder)
library(gapminder)Take a look at the gapminder data frame that is now available to you:
gapminder# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Afghanistan Asia 1952 28.8 8425333 779.
2 Afghanistan Asia 1957 30.3 9240934 821.
3 Afghanistan Asia 1962 32.0 10267083 853.
4 Afghanistan Asia 1967 34.0 11537966 836.
5 Afghanistan Asia 1972 36.1 13079460 740.
6 Afghanistan Asia 1977 38.4 14880372 786.
7 Afghanistan Asia 1982 39.9 12881816 978.
8 Afghanistan Asia 1987 40.8 13867957 852.
9 Afghanistan Asia 1992 41.7 16317921 649.
10 Afghanistan Asia 1997 41.8 22227415 635.
# ℹ 1,694 more rowsThe first 10 rows are printed, with the following columns:
| column name | data type | meaning |
|---|---|---|
country |
factor (<fct>) |
Country |
continent |
factor (<fct>) |
Continent that the country is in |
year |
integer (<int>) |
Focal year for the stats in the next columns |
lifeExp |
integer (<int>) |
Mean life expectancy in years |
pop |
integer (<int>) |
Population size |
gdpPercap |
double (<dbl>) |
Per-capita GDP |
Two columns are of type factor, an alternative to character commonly used for categorical data.
You can also use the View() function to look at the data frame. This will open a new tab in your editor pane with a spreadsheet-like look and feel:
View(gapminder)
# (Should display the dataset in an editor pane tab)In the head() output above, the gapminder data frame is referred to as a “tibble”, which is just a slight variation on data frames from the tidyverse4.
3 arrange()
The arrange() function is like sorting functionality in Excel: it changes the order of rows based on the values in one or more columns. It will always rearrange the order of rows as a whole, never just of individual columns since that would scramble the data.
gapminder is currently first sorted alphabetically by country, and next by year. But you may, for example, want to sort by population size instead:
gm_arranged <- arrange(.data = gapminder, pop)Before we take a look at the result, let’s break down the syntax of the arrange() function:
- The first argument is the data frame to be sorted, which we specify with the
.dataargument. - The second argument is the column to sort by, which we specify without an argument name.
Also, like all dplyr functions, arrange() outputs a new data frame, and does not modify the input data frame. In this case, we chose to assign the output to a new object called gm_arranged.
What would have happened if we had assigned the output to gapminder?
Now let’s look at the result of the sorting:
gm_arranged# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Sao Tome and Principe Africa 1952 46.5 60011 880.
2 Sao Tome and Principe Africa 1957 48.9 61325 861.
3 Djibouti Africa 1952 34.8 63149 2670.
4 Sao Tome and Principe Africa 1962 51.9 65345 1072.
5 Sao Tome and Principe Africa 1967 54.4 70787 1385.
6 Djibouti Africa 1957 37.3 71851 2865.
7 Sao Tome and Principe Africa 1972 56.5 76595 1533.
8 Sao Tome and Principe Africa 1977 58.6 86796 1738.
9 Djibouti Africa 1962 39.7 89898 3021.
10 Sao Tome and Principe Africa 1982 60.4 98593 1890.
# ℹ 1,694 more rowsSorting can help you find observations with the smallest or largest values for a certain column: above, we see that the smallest population size in the dataset is Sao Tome and Principe in 1952.
Default sorting is from small to large, as seen above . To sort in reverse order, use the desc() (descending) helper function:
arrange(.data = gapminder, desc(pop))# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 China Asia 2007 73.0 1318683096 4959.
2 China Asia 2002 72.0 1280400000 3119.
3 China Asia 1997 70.4 1230075000 2289.
4 China Asia 1992 68.7 1164970000 1656.
5 India Asia 2007 64.7 1110396331 2452.
6 China Asia 1987 67.3 1084035000 1379.
7 India Asia 2002 62.9 1034172547 1747.
8 China Asia 1982 65.5 1000281000 962.
9 India Asia 1997 61.8 959000000 1459.
10 China Asia 1977 64.0 943455000 741.
# ℹ 1,694 more rowsHere, we didn’t assign the output to a new object, so it was just printed to screen. We’ll keep doing that for the rest of the examples in this session, so we can more easily see the results of the operations we are performing.
Finally, you may want to sort by multiple columns, where ties in the first column are broken by a second column (and so on). Do this by simply listing the columns in the appropriate order:
# Sort first by continent, then by country:
arrange(.data = gapminder, continent, country)# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Algeria Africa 1952 43.1 9279525 2449.
2 Algeria Africa 1957 45.7 10270856 3014.
3 Algeria Africa 1962 48.3 11000948 2551.
4 Algeria Africa 1967 51.4 12760499 3247.
5 Algeria Africa 1972 54.5 14760787 4183.
6 Algeria Africa 1977 58.0 17152804 4910.
7 Algeria Africa 1982 61.4 20033753 5745.
8 Algeria Africa 1987 65.8 23254956 5681.
9 Algeria Africa 1992 67.7 26298373 5023.
10 Algeria Africa 1997 69.2 29072015 4797.
# ℹ 1,694 more rowsPay attention to the syntax: additional columns to sort by are additional arguments to the function, not part of the same argument. Other dplyr functions work equivalently in this regard.
Challenge 1: arrange()
Which line of code would you use to find out which observation (country and year) has the highest per-capita GDP in the entire dataset?
# A)
arrange(.data = gapminder, desc(gdpPercap), year, country)
# B)
arrange(.data = gapminder, country, year, gdpPercap)
# C)
arrange(.data = gapminder, desc(gdpPercap))
# D)
arrange(.data = gapminder, gdpPercap)4 filter()
The filter() function outputs only those rows that satisfy one or more conditions. It is similar to Filter functionality in Excel.
4.1 Filter based on one condition
This first filter() example outputs only rows for which the life expectancy exceeds 80 years (remember, each row represents a country in a given year):
filter(.data = gapminder, lifeExp > 80)# A tibble: 21 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Australia Oceania 2002 80.4 19546792 30688.
2 Australia Oceania 2007 81.2 20434176 34435.
3 Canada Americas 2007 80.7 33390141 36319.
4 France Europe 2007 80.7 61083916 30470.
5 Hong Kong, China Asia 2002 81.5 6762476 30209.
6 Hong Kong, China Asia 2007 82.2 6980412 39725.
7 Iceland Europe 2002 80.5 288030 31163.
8 Iceland Europe 2007 81.8 301931 36181.
9 Israel Asia 2007 80.7 6426679 25523.
10 Italy Europe 2002 80.2 57926999 27968.
# ℹ 11 more rowsHow many rows were output?
As the example above demonstrated, filter() outputs rows that satisfy the condition(s) you specify. These conditions don’t have to be based on numeric comparisons:
# Only keep rows where the value in the 'continent' column is 'Europe':
filter(.data = gapminder, continent == "Europe")# A tibble: 360 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Albania Europe 1952 55.2 1282697 1601.
2 Albania Europe 1957 59.3 1476505 1942.
3 Albania Europe 1962 64.8 1728137 2313.
4 Albania Europe 1967 66.2 1984060 2760.
5 Albania Europe 1972 67.7 2263554 3313.
6 Albania Europe 1977 68.9 2509048 3533.
7 Albania Europe 1982 70.4 2780097 3631.
8 Albania Europe 1987 72 3075321 3739.
9 Albania Europe 1992 71.6 3326498 2497.
10 Albania Europe 1997 73.0 3428038 3193.
# ℹ 350 more rowsRemember to use two equals signs == to test for equality!
4.2 Filter based on multiple conditions
It’s also possible to filter based on multiple conditions. For example, you may want to see which countries in Asia had a life expectancy greater than 80 years:
filter(.data = gapminder, continent == "Asia", lifeExp > 80)# A tibble: 6 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Hong Kong, China Asia 2002 81.5 6762476 30209.
2 Hong Kong, China Asia 2007 82.2 6980412 39725.
3 Israel Asia 2007 80.7 6426679 25523.
4 Japan Asia 1997 80.7 125956499 28817.
5 Japan Asia 2002 82 127065841 28605.
6 Japan Asia 2007 82.6 127467972 31656.Like above, by default, multiple conditions are combined using a Boolean AND. In other words, in a given row, each condition must be met to output the row.
If you want to combine conditions using a Boolean OR, where only one of the conditions needs to be met, use a | (vertical bar) between the conditions:
# Keep rows with a high life expectancy and/or a high GDP:
filter(.data = gapminder, lifeExp > 80 | gdpPercap > 100000)# A tibble: 24 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Australia Oceania 2002 80.4 19546792 30688.
2 Australia Oceania 2007 81.2 20434176 34435.
3 Canada Americas 2007 80.7 33390141 36319.
4 France Europe 2007 80.7 61083916 30470.
5 Hong Kong, China Asia 2002 81.5 6762476 30209.
6 Hong Kong, China Asia 2007 82.2 6980412 39725.
7 Iceland Europe 2002 80.5 288030 31163.
8 Iceland Europe 2007 81.8 301931 36181.
9 Israel Asia 2007 80.7 6426679 25523.
10 Italy Europe 2002 80.2 57926999 27968.
# ℹ 14 more rows5 The pipe (|>)
Examples so far applied a single dplyr function to a data frame. But in practice, it’s common to use several consecutive dplyr functions to wrangle a dataframe into the format you want.
For example, you may want to filter rows and then sort the result. You could do that as follows:
gm_filt <- filter(.data = gapminder, lifeExp < 50, year > 2000)
arrange(.data = gm_filt, desc(pop))# A tibble: 42 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Nigeria Africa 2007 46.9 135031164 2014.
2 Nigeria Africa 2002 46.6 119901274 1615.
3 Congo, Dem. Rep. Africa 2007 46.5 64606759 278.
4 Congo, Dem. Rep. Africa 2002 45.0 55379852 241.
5 South Africa Africa 2007 49.3 43997828 9270.
6 Tanzania Africa 2002 49.7 34593779 899.
7 Afghanistan Asia 2007 43.8 31889923 975.
8 Afghanistan Asia 2002 42.1 25268405 727.
9 Uganda Africa 2002 47.8 24739869 928.
10 Mozambique Africa 2007 42.1 19951656 824.
# ℹ 32 more rowsAnd you could go on along these lines, successively creating new objects that you then use for the next step.
But there is a more elegent way of dong this, directly sending (“piping”) output from one function into the next function with the pipe operator |> (a vertical bar followed by a greater-than sign). Let’s start by seeing a reformulation of the code above with pipes:
gapminder |>
filter(lifeExp < 50, year > 2000) |>
arrange(desc(pop))# A tibble: 42 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Nigeria Africa 2007 46.9 135031164 2014.
2 Nigeria Africa 2002 46.6 119901274 1615.
3 Congo, Dem. Rep. Africa 2007 46.5 64606759 278.
4 Congo, Dem. Rep. Africa 2002 45.0 55379852 241.
5 South Africa Africa 2007 49.3 43997828 9270.
6 Tanzania Africa 2002 49.7 34593779 899.
7 Afghanistan Asia 2007 43.8 31889923 975.
8 Afghanistan Asia 2002 42.1 25268405 727.
9 Uganda Africa 2002 47.8 24739869 928.
10 Mozambique Africa 2007 42.1 19951656 824.
# ℹ 32 more rowsWhat happened here? We took the gapminder data frame, sent (“piped”) it into the filter() function, whose output in turn was piped into the arrange() function. You can think of the pipe as “then”: take gapminder, then filter, then arrange.
When using the pipe, you no longer specify the input data frame with the .data argument, because the function now gets its input data via the pipe (specifically, the input goes to the function’s first argument by default).
Using pipes involves less typing and is especially more readable than using successive assignments5. For code readability, it is good practice to always start a new line after a pipe |>, and to keep the subsequent line(s) indented as RStudio will automatically do.
Challenge 2: Find the mistakes
This pipeline is supposed to find the 3 countries with the lowest GDP per capita in 1997, sorted from lowest to highest. It has two errors. Find and fix them:
gapminder
filter(year = 1997) |>
arrange(gdpPercap) |>
head(n = 3)Click for the solution
- Line 1: Missing pipe
|>after gapminder - Line 2: Should use
==(not=) for comparison
Corrected code:
gapminder |>
filter(year == 1997) |>
arrange(gdpPercap) |>
head(n = 3)# A tibble: 3 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Congo, Dem. Rep. Africa 1997 42.6 47798986 312.
2 Myanmar Asia 1997 60.3 43247867 415
3 Burundi Africa 1997 45.3 6121610 463.Challenge 3: Find the extremes
Using filter(), arrange(), and the pipe, answer:
Which African country had the highest life expectancy in 2002?
Click for the solution
(We’ll use the same trick with
head()like in the exercise above to only see the top 1 country, but this is not strictly necessary.)gapminder |> filter(continent == "Africa", year == 2002) |> arrange(desc(lifeExp)) |> head(n = 1)# A tibble: 1 × 6 country continent year lifeExp pop gdpPercap <fct> <fct> <int> <dbl> <int> <dbl> 1 Reunion Africa 2002 75.7 743981 6316.What was the smallest population recorded in Asia across all years?
Click for the solution
(We’ll use the same trick with
head()like in the exercise above to only see the bottom 1 country, but this is not strictly necessary.)gapminder |> filter(continent == "Asia") |> arrange(pop) |> head(n = 1)# A tibble: 1 × 6 country continent year lifeExp pop gdpPercap <fct> <fct> <int> <dbl> <int> <dbl> 1 Bahrain Asia 1952 50.9 120447 9867.
6 mutate()
So far, we’ve focused on functions that subset and reorganize data frames. But you haven’t seen how you can change the data or compute derived data.
This can be done with the mutate() function. For example, to create a new column with population sizes in millions:
# Create a new column 'pop_million' by dividing 'pop' by a million:
gapminder |>
mutate(pop_million = pop / 10^6)# A tibble: 1,704 × 7
country continent year lifeExp pop gdpPercap pop_million
<fct> <fct> <int> <dbl> <int> <dbl> <dbl>
1 Afghanistan Asia 1952 28.8 8425333 779. 8.43
2 Afghanistan Asia 1957 30.3 9240934 821. 9.24
3 Afghanistan Asia 1962 32.0 10267083 853. 10.3
4 Afghanistan Asia 1967 34.0 11537966 836. 11.5
5 Afghanistan Asia 1972 36.1 13079460 740. 13.1
6 Afghanistan Asia 1977 38.4 14880372 786. 14.9
7 Afghanistan Asia 1982 39.9 12881816 978. 12.9
8 Afghanistan Asia 1987 40.8 13867957 852. 13.9
9 Afghanistan Asia 1992 41.7 16317921 649. 16.3
10 Afghanistan Asia 1997 41.8 22227415 635. 22.2
# ℹ 1,694 more rowsTo modify an existing column rather than adding a new one, simply “assign back to the same name”:
# Change the unit of the 'pop' column:
gapminder |>
mutate(pop = pop / 10^6)# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <dbl> <dbl>
1 Afghanistan Asia 1952 28.8 8.43 779.
2 Afghanistan Asia 1957 30.3 9.24 821.
3 Afghanistan Asia 1962 32.0 10.3 853.
4 Afghanistan Asia 1967 34.0 11.5 836.
5 Afghanistan Asia 1972 36.1 13.1 740.
6 Afghanistan Asia 1977 38.4 14.9 786.
7 Afghanistan Asia 1982 39.9 12.9 978.
8 Afghanistan Asia 1987 40.8 13.9 852.
9 Afghanistan Asia 1992 41.7 16.3 649.
10 Afghanistan Asia 1997 41.8 22.2 635.
# ℹ 1,694 more rowsChallenge 4: Absolute GDP
Use mutate() to create a new column gdp_billion that has the absolute GDP (i.e., not relative to population size) in units of billions.
Click for the solution
You need to do two things, which can be combined into a single line:
- Make the GDP absolute by multiplying by the population size:
gdpPercap * pop - Change the unit of the absolute GDP to billions by dividing by a billion:
/ 10^9
gapminder |>
mutate(gdp_billion = gdpPercap * pop / 10^9)# A tibble: 1,704 × 7
country continent year lifeExp pop gdpPercap gdp_billion
<fct> <fct> <int> <dbl> <int> <dbl> <dbl>
1 Afghanistan Asia 1952 28.8 8425333 779. 6.57
2 Afghanistan Asia 1957 30.3 9240934 821. 7.59
3 Afghanistan Asia 1962 32.0 10267083 853. 8.76
4 Afghanistan Asia 1967 34.0 11537966 836. 9.65
5 Afghanistan Asia 1972 36.1 13079460 740. 9.68
6 Afghanistan Asia 1977 38.4 14880372 786. 11.7
7 Afghanistan Asia 1982 39.9 12881816 978. 12.6
8 Afghanistan Asia 1987 40.8 13867957 852. 11.8
9 Afghanistan Asia 1992 41.7 16317921 649. 10.6
10 Afghanistan Asia 1997 41.8 22227415 635. 14.1
# ℹ 1,694 more rows7 summarize()
The final function dplyr function we’ll cover is summarize(), which computes summaries of your data across rows. For example, to calculate the mean GDP across the entire dataset:
# The syntax is similar to 'mutate': <new-column> = <operation>
gapminder |>
summarize(mean_gdp = mean(gdpPercap))# A tibble: 1 × 1
mean_gdp
<dbl>
1 7215.The output is still a dataframe, but unlike with all previous dplyr functions, it is completely different from the input dataframe, “collapsing” the data down to as little as a single number, like above.
summarize() becomes really powerful in combination with the helper function group_by() to compute groupwise stats. For example, to get the mean GDP separately for each continent:
gapminder |>
group_by(continent) |>
summarize(mean_gdp = mean(gdpPercap))# A tibble: 5 × 2
continent mean_gdp
<fct> <dbl>
1 Africa 2194.
2 Americas 7136.
3 Asia 7902.
4 Europe 14469.
5 Oceania 18622.group_by() implicitly splits a data frame into groups of rows: here, one group for observations from each continent. After that, operations like in summarize() will happen separately for each group, which is how we ended up with per-continent means.
Challenge 5: Mean life expectancy
Calculate the average life expectancy per country. Which country has the longest average life expectancy and which has the shortest average life expectancy?
Click for some hints
Since you’ll need to examine your output in two ways, it makes sense to store the output of the life expectancy calculation in a dataframe.
Next, an easy way to get rows with lowest and highest values in a column is to simply sort by that column and examining the output.
Click for the solution
First, create a dataframe with the mean life expectancy by country:
lifeExp_bycountry <- gapminder |>
group_by(country) |>
summarize(mean_lifeExp = mean(lifeExp))Then, arrange that dataframe in two directions to see the countries with the longest and shortest life expectancy. You could optionally pipe into head() to only see the top n, here top 1:
lifeExp_bycountry |>
arrange(mean_lifeExp) |>
head(n = 1)# A tibble: 1 × 2
country mean_lifeExp
<fct> <dbl>
1 Sierra Leone 36.8lifeExp_bycountry |>
arrange(desc(mean_lifeExp)) |>
head(n = 1)# A tibble: 1 × 2
country mean_lifeExp
<fct> <dbl>
1 Iceland 76.58 Bonus material for self-study
8.1 Writing and reading tabular data to and from files
When working with your own data in R, you’ll usually need to read data from files into your R environment, and conversely, to write data that is in your R environment to files.
While it’s possible to make R interact with Excel spreadsheet files6, storing your data in plain-text files generally benefits reproducibility. Tabular plain text files can be stored using:
- A Tab as the column delimiter (often called TSV files, and stored with a
.tsvextension) - A comma as the column delimiter (often called CSV files, and stored with a
.csvextension).
To practice writing and reading data to and from TSV files, the examples below use functions from the readr package. This package is part of the core tidyverse and therefore already loaded into your environment.
Writing to files
To write the gapminder data frame to a TSV file, use the write_tsv() function with argument x for the R object and argument file for the file path:
write_tsv(x = gapminder, file = "gapminder.tsv")Because we simply provided a file name, the file will have been written in your R working directory7.
If you want to take a look at the file you just created, you can find it in RStudio’s Files tab and click on it, which will open it in the editor panel. Of course, you could also find it in your computer’s file browser and open it in different ways.
The file argument to write_tsv() takes a file path, meaning that you can specify any location on your computer for it, in addition to the file name.
For example, if you had a folder called results in your current working directory, you could save the file in there like so:
write_tsv(x = gapminder, file = "results/gapminder.tsv")Note that a forward slash / as a folder delimiter will work regardless of your operating system (even though Windows natively delimits folder by a backslash \).
Finally, if you want to store the file in a totally different place than your working dir, it’s good to know that you can also use a so-called absolute (or “full”) path. For example:
write_tsv(x = gapminder, file = "/Users/poelstra.1/Desktop/gapminder.tsv")Reading from files
To practice reading data from a file, use the read_tsv() function on the file you just created:
gapminder_reread <- read_tsv(file = "gapminder.tsv")Rows: 1704 Columns: 6
── Column specification ────────────────────────────────────────────────────────
Delimiter: "\t"
chr (2): country, continent
dbl (4): year, lifeExp, pop, gdpPercap
ℹ Use `spec()` to retrieve the full column specification for this data.
ℹ Specify the column types or set `show_col_types = FALSE` to quiet this message.Note that this function is rather chatty, telling you how many rows and columns it read, and what their data types are. Let’s check the resulting object:
gapminder_reread# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<chr> <chr> <dbl> <dbl> <dbl> <dbl>
1 Afghanistan Asia 1952 28.8 8425333 779.
2 Afghanistan Asia 1957 30.3 9240934 821.
3 Afghanistan Asia 1962 32.0 10267083 853.
4 Afghanistan Asia 1967 34.0 11537966 836.
5 Afghanistan Asia 1972 36.1 13079460 740.
6 Afghanistan Asia 1977 38.4 14880372 786.
7 Afghanistan Asia 1982 39.9 12881816 978.
8 Afghanistan Asia 1987 40.8 13867957 852.
9 Afghanistan Asia 1992 41.7 16317921 649.
10 Afghanistan Asia 1997 41.8 22227415 635.
# ℹ 1,694 more rowsThis looks good! Do note that the column’s data types are not identical to what they were (year and pop are saved as double rather than integer, and country and continent as character rather than factor). This is largely expected because that kind of metadata is not stored in a plain-text TSV, so read_tsv() will by default simply make best guesses as to the types.
Alternatively, you could tell read_tsv() what the column types should be, using the col_types argument (with abbreviations f for factor, i for integer, d for double — run ?read_tsv for more info):
read_tsv(file = "gapminder.tsv", col_types = "ffidi")# A tibble: 1,704 × 6
country continent year lifeExp pop gdpPercap
<fct> <fct> <int> <dbl> <int> <dbl>
1 Afghanistan Asia 1952 28.8 8425333 779.
2 Afghanistan Asia 1957 30.3 9240934 821.
3 Afghanistan Asia 1962 32.0 10267083 853.
4 Afghanistan Asia 1967 34.0 11537966 836.
5 Afghanistan Asia 1972 36.1 13079460 740.
6 Afghanistan Asia 1977 38.4 14880372 786.
7 Afghanistan Asia 1982 39.9 12881816 978.
8 Afghanistan Asia 1987 40.8 13867957 852.
9 Afghanistan Asia 1992 41.7 16317921 649.
10 Afghanistan Asia 1997 41.8 22227415 635.
# ℹ 1,694 more rowsTo read and write CSV files instead, use read_csv() / write_csv() in the same way.
8.2 rename() to change column names
The rename() dplyr function is one of the package’s simplest, but it is very useful when you want to change column names.
The syntax to specify the new and old name within the function is new_name = old_name. For example, to rename the gdpPercap column:
rename(.data = gapminder, gdp_per_capita = gdpPercap)# A tibble: 1,704 × 6
country continent year lifeExp pop gdp_per_capita
<fct> <fct> <int> <dbl> <int> <dbl>
1 Afghanistan Asia 1952 28.8 8425333 779.
2 Afghanistan Asia 1957 30.3 9240934 821.
3 Afghanistan Asia 1962 32.0 10267083 853.
4 Afghanistan Asia 1967 34.0 11537966 836.
5 Afghanistan Asia 1972 36.1 13079460 740.
6 Afghanistan Asia 1977 38.4 14880372 786.
7 Afghanistan Asia 1982 39.9 12881816 978.
8 Afghanistan Asia 1987 40.8 13867957 852.
9 Afghanistan Asia 1992 41.7 16317921 649.
10 Afghanistan Asia 1997 41.8 22227415 635.
# ℹ 1,694 more rows8.3 select() to pick columns (variables)
The select() dplyr function subsets a data frame by including/excluding certain columns. By default, it only includes the columns you specify:
gapminder |>
select(year, country, pop)# A tibble: 1,704 × 3
year country pop
<int> <fct> <int>
1 1952 Afghanistan 8425333
2 1957 Afghanistan 9240934
3 1962 Afghanistan 10267083
4 1967 Afghanistan 11537966
5 1972 Afghanistan 13079460
6 1977 Afghanistan 14880372
7 1982 Afghanistan 12881816
8 1987 Afghanistan 13867957
9 1992 Afghanistan 16317921
10 1997 Afghanistan 22227415
# ℹ 1,694 more rowsThe order of the columns in the output data frame is exactly as you list them in select(), and doesn’t need to be the same as in the input data frame. In other words, select() is also one way to reorder columns: in the example above, we made year appear before country.
You can also specify columns that should be excluded, by prefacing their name with a ! (or a -):
# This will include all columns _except_ continent:
gapminder |>
select(!continent)# A tibble: 1,704 × 5
country year lifeExp pop gdpPercap
<fct> <int> <dbl> <int> <dbl>
1 Afghanistan 1952 28.8 8425333 779.
2 Afghanistan 1957 30.3 9240934 821.
3 Afghanistan 1962 32.0 10267083 853.
4 Afghanistan 1967 34.0 11537966 836.
5 Afghanistan 1972 36.1 13079460 740.
6 Afghanistan 1977 38.4 14880372 786.
7 Afghanistan 1982 39.9 12881816 978.
8 Afghanistan 1987 40.8 13867957 852.
9 Afghanistan 1992 41.7 16317921 649.
10 Afghanistan 1997 41.8 22227415 635.
# ℹ 1,694 more rowsThere are also ways to e.g. select ranges of columns that are beyond the scope of this short workshop. Check the select() help by typing ?select to learn more.
8.4 Counting rows with count()
A common operation is simply counting the number of observations for each group. For instance, to check the number of countries included in the dataset for the year 2002, you can use the count() function. It takes the name of one or more columns to define the groups we are interested in, and can optionally sort the results in descending order with sort = TRUE:
gapminder |>
filter(year == 2002) |>
count(continent, sort = TRUE)# A tibble: 5 × 2
continent n
<fct> <int>
1 Africa 52
2 Asia 33
3 Europe 30
4 Americas 25
5 Oceania 28.5 Function name conflicts
When you loaded the tidyverse, the output included a “Conflicts” section that may have seemed ominous:
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorsWhat this means is that two tidyverse functions, filter() and lag(), have the same names as two functions from the stats package that were already in your R environment.
Those stats package functions are part of what is often referred to as “base R”: core R functionality that is always available (loaded) when you start R.
Due to this function name conflict/collision, the filter() function from dplyr “masks” the filter() function from stats. Therefore, if you write a command with filter(), it will use the dplyr function and not the stats function.
You can use a “masked” function, by prefacing it with its package name as follows: stats::filter().
8.6 Using the pipe keyboard shortcut
The following RStudio keyboard shortcut will insert a pipe symbol: Ctrl/Cmd + Shift + M.
However, by default this (still) inserts an older pipe symbol, %>%. As long as you’ve loaded the tidyverse8, it would not really be a problem to just use %>% instead of |>. But it would be preferred to make RStudio insert the |> pipe symbol:
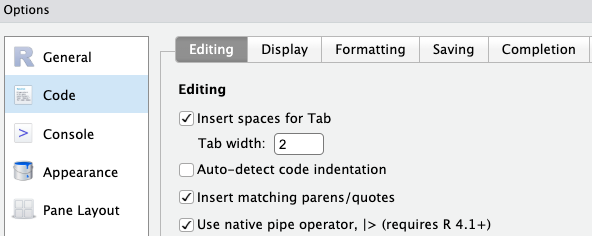
- Click
Tools>Global Options>Codetab on the right - Check the box “Use native pipe operator” as shown below
- Click
OKat the bottom of the options dialog box to apply the change and close the box.
8.7 Learn even more
The material in this page was adapted from this Carpentries lesson episode, which has a lot more content than we were able to cover today.
Regarding data wrangling specifically, here are two particularly useful additional skills to learn:
Pivoting/reshaping — moving between ‘wide’ and ‘long’ data formats with
pivot_wider()andpivot_longer(). This is covered in episode 13 of the focal Carpentries lesson.Joining/merging — combining multiple dataframes based on one or more shared columns. This can be done with dplyr’s
join_*()functions – see for example this chapter in R for Data Science.
Footnotes
Or Click
File=>New file=>R Script.↩︎If not: run
install.packages("tidyverse")now.↩︎If not: run
install.packages("gapminder")now.↩︎The main difference is the nicer default printing behavior of tibbles: e.g. the data types of columns are shown, and only a limited number of rows are printed.↩︎
Using pipes is also faster and uses less computer memory.↩︎
Using the readxl package. This is installed as part of the tidyverse but is not a core tidyverse package and therefore needs to be loaded separately:
library(readxl).↩︎Recall that you can see where that is at the top of the Console tab, or by running
getwd().↩︎This symbol only works when you have one of the tidyverse packages loaded.↩︎